Fig. 1
Download original image
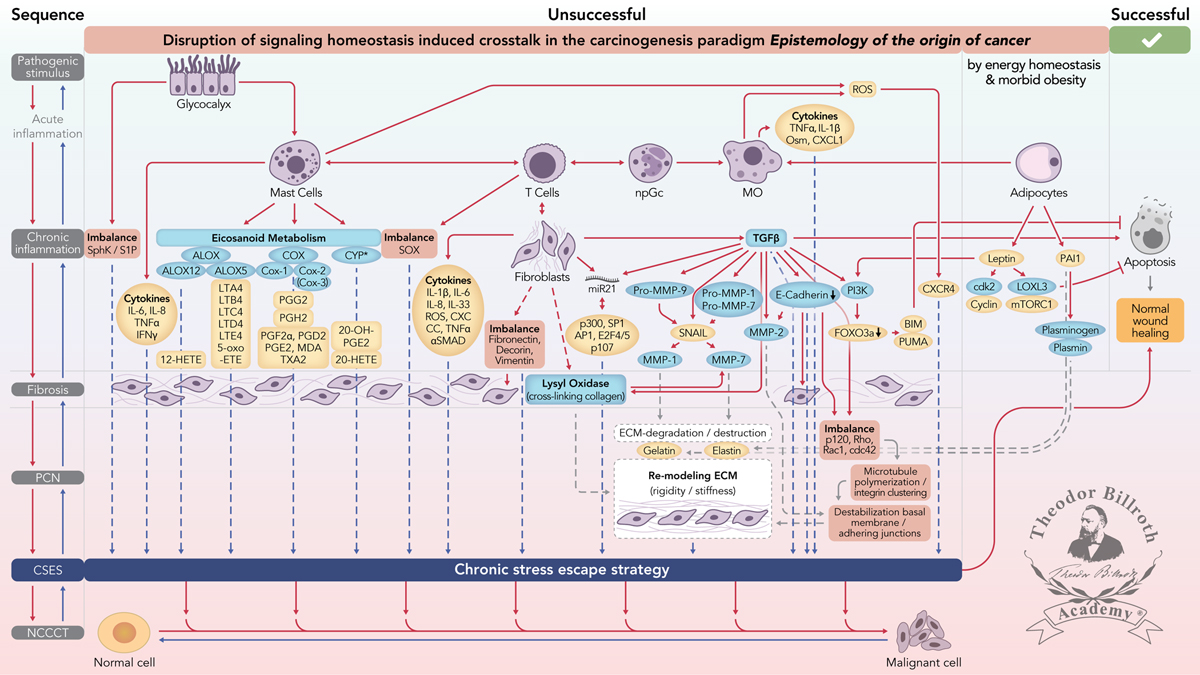
Simplified scheme of the disruption of signaling homeostasis-induced crosstalk in the carcinogenesis paradigm “epistemology of the origin of cancer” consisting of a six-step sequence: (1) a pathogenic stimulus followed by (2) chronic inflammation from which develops (3) fibrosis with associated remodeling of the cellular microenvironment; and from these changes a (4) precancerous niche (PCN), a product of fibrosis, with remodeling by persistent inflammation, develops that triggers the deployment of (5) a chronic stress escape strategy and when this fails resolve it by (6) normal cell to cancerous cell transition (NCCCT) by PCN-induced cell matrix stress occurs. This figure was published as original illustration in paper 3 of this Special Issue – Disruption of homeostasis-induced signaling and crosstalk in the carcinogenesis paradigm “Epistemology of the origin of cancer” entitled “Chronic inflammation evoked by pathogenic stimulus during carcinogenesis”. We point out, that to the complexity of the content of the Special Issue the original and/or modified version of the original illustration was republished within the following papers of the Special Issue: paper 5 “Microbiome and morbid obesity increase pathogenic stimulus diversity”, paper 6 “Precancerous niche (PCN), a product of fibrosis with remodeling by incessant chronic inflammation”, paper 7 “Metformin alters signaling homeostasis”, paper 8 “Transition from normal to cancerous cell by precancerous niche (PCN) induced chronic cell-matrix stress” and paper 9 “NF-κB signaling and crosstalk during carcinogenesis”. Nomenclature: The nomenclature common abbreviations are in bold, followed by the common trivial names (if available) and (if available) by the name in accordance to the International Union of Pure and Applied Chemistry (IUPAC): PCN precancerous niche; CSES chronic stress escape strategy; NCCCT normal cell to cancerous cell transition; SphK sphingosine kinase isoform; S1P sphingosine-1-phosphate; IL-6 interleukin 6; IL-8 interleukin 8; TNFα tumor necrosis factor alpha; IFNγ interferon gamma; ALOX lipoxygenase, arachidonate lipoxygenase; ALOX12 12-lipoxygenase, 12-LOX, 12S-LOX, arachidonate 12-lipoxygenase 12S type; ALOX5 5-lipoxygenase, 5-LOX, arachidonate 5-lipoxygenase; 12-HETE 12-hydroxyeicosatetraenoic acid; LTA4 leukotriene A4, 4-[(2S,3S)-3-[(1E,3E,5Z,8Z)-tetradeca-1,3,5,8-tetraenyl]oxiran-2-yl]butanoic acid; LTB4 leukotriene B4, (5S,6Z,8E,10E,12R,14Z)-5,12-dihydroxyicosa-6,8,10,14-tetraenoic acid; LTC4 leukotriene C4, (5S,6R,7E,9E,11Z,14Z)-6-[(2R)-2-[[(4S)-4-amino-4-carboxybutanoyl]amino]-3-(carboxymethylamino)-3-oxopropyl]sulfanyl-5-hydroxyicosa-7,9,11,14-tetraenoic acid; LTD4 leukotriene D4, (5S,6R,7E,9E,11Z,14Z)-6-[(2R)-2-amino-3-(carboxymethylamino)-3-oxopropyl]sulfanyl-5-hydroxyicosa-7,9,11,14-tetraenoic acid; LTE4 leukotriene E4, (5S,6R,7E,9E,11Z,14Z)-6-[(2R)-2-amino-2-carboxyethyl]sulfanyl-5-hydroxyicosa-7,9,11,14-tetraenoic acid; 5-oxo-ETE (6E,8Z,11Z,14Z)-5-oxoicosa-6,8,11,14-tetraenoic acid; Cox cyclooxygenase; Cox-1 cyclooxygenase 1; Cox-2 cyclooxygenase 2; Cox-3 isoform of Cox-2 (therefore in brakes); PGG2 prostaglandin G2, (Z)-7-[(1S,4R,5R,6R)-5-[(E,3S)-3-hydroperoxyoct-1-enyl]-2,3-dioxabicyclo[2.2.1]heptan-6-yl]hept-5-enoic acid; PGH2 prostaglandin H2, (Z)-7-[(1S,4R,5R,6R)-5-[(E,3S)-3-hydroxyoct-1-enyl]-2,3-dioxabicyclo[2.2.1]heptan-6-yl]hept-5-enoic acid; PGFF2α prostaglandine F2 alpha, (Z)-7-[(1R,2R,3R,5S)-3,5-dihydroxy-2-[(E,3S)-3-hydroxyoct-1-enyl]cyclopentyl]hept-5-enoic acid; PGD2 prostaglandin D2, (Z)-7-[(1R,2R,5S)-5-hydroxy-2-[(E,3S)-3-hydroxyoct-1-enyl]-3-oxocyclopentyl]hept-5-enoic acid; PGE2 prostaglandin E2, (Z)-7-[(1R,2R,3R)-3-hydroxy-2-[(E,3S)-3-hydroxyoct-1-enyl]-5-oxocyclopentyl]hept-5-enoic acid; MDA malondialdehyde, propanedial; TXA2 thromboxane A2, (Z)-7-[(1S,2S,3R,5S)-3-[(E,3S)-3-hydroxyoct-1-enyl]-4,6-dioxabicyclo[3.1.1]heptan-2-yl]hept-5-enoic acid; CYP* cytochrome P450 isoforms; 20-OH-PGE2 20-hydroxy prostaglandin E2; 20-HETE 20-hydroxyeicosatetraenoic acid, (5Z,8Z,11Z,14Z)-20-hydroxyicosa-5,8,11,14-tetraenoic acid; SOX [sex-determining region Y (Sry) box-containing] transcription factor family; IL-β1 interleukin beta 1; IL-33 interleukin 33; ROS reactive oxygen species; CXC CC chemokine receptors; αSMAD alpha-smooth muscle actin; miR21 micro RNA-21; p300 protein 300 (p300-CBP coactivator family); SP1 specificity protein 1; AP1 activator protein 1; E2F4/5 cytoplasmic complex of Smad3, retinoblastoma-like protein 1 (P107, RBL1), E2F4/5 and d-prostanoid (DP1); p107 retinoblastoma-like protein 1, RBL1; TGFβ transforming growth factor beta; Pro-MMP-9 pro-matrix metalloproteinase 9; Pro-MMP-1 pro-matrix metalloproteinase 1; Pro-MMP-7 pro-matrix metalloproteinase 7; SNAIL zinc finger protein SNAI1; MMP-1 matrix metalloproteinase 1; MMP-7 matrix metalloproteinase 7; MMP-2 matrix metalloproteinase 2; E-Cadherin CAM 120/80 or epithelial cadherin, cadherin-1, epithelial cadherin; CXCL1 chemokine (C–X–C motif) ligand 1; Osm oncostatin-M; PI3K phosphatidylinositide 3-kinase; FOXO3a forkhead box protein O3a; p120 catenin delta-1, protein 120; Rho Ras homolog gene family, member A; Rac1 Ras-related C3 botulinum toxin substrate 1; cdc42 cell division control protein 42 homolog; BIM Bcl-2 interacting mediator of cell death; PUMA BH3-only protein; CXCR4 C–X–C motif of chemokine receptor 4; cdk2 cyclin-dependent kinase 2;LOXL3 lysyl oxidase homolog 3; mTORc1 rapamycin complex 1; PAI1 plasminogen activator inhibitor-1.

